Simulation and Bootstrapping
Simulation
Many people think of a coin toss when they think about
randomness. While
everything in nature is random, a coin toss has the extra quality
of being a completely known process. We do not know what the
outcome of the next coin is going to be, but we know the average
behavior quite well, viz., the outcome will be either a
Head or a Tail, each of which is equally likely. It is this
perfect knowledge that leads us to use a coin toss when making a
fair decision.
A coin toss is a random mechanism that we can repeat at our will
to produce random quantities with a known behavior. Another such
a device is a die. But such practical devices used to be limited in
number and scope before the advent of high speed computers.
A remarkable fact about computers is that they can be instructed
to produce random quantities whose general behavior is
completely under control. We can, for example make the computer
toss a coin
with exactly a specified amount of bias for Head! We can turn the
computer easily into a 100-faced die, something that is almost to
construct in practice. In fact, computers can be made to perform
more exotic (as well more useful) random computations that have
started to play a dominant role in statistics.
In this last tutorial we shall take a look at these.
Putting it in perspective
Man has devised various techniques to deal with chance. These may
be classified into roughly three categories:
-
Wait and see: Here we patiently wait until the end of the
activity. This time-honored method
remains the most frequently used one even today.
Who can deny the usefulness of observing the sky carefully to learn about the random patterns?
However, learning by just trial-and-error may
prove too expensive. When we want to design a space station to
maximize its chance of capturing some rare phenomenon, it would
prove it bit too costly to simply rely on trial and error!
-
Use probability models: Often we have enough information
(theoretical and/or based on prior observations)
about the mechanism underlying a random phenomenon.
For example, if a careful scrutiny of a die does not
reveal any difference between the faces of a die, one may assume
that the die is fair. Such information can be expressed as a
probability distribution:
The outcome X can take values 1,...,6 each with equal
probability (taken to be 1/6 to keep the sum 1).
Using this mathematical formulation of our
knowledge about the random phenomenon we can now analyze the
distribution mathematically. For example, consider the following
gambling game
A die is rolled, and the casino owner gives you Re 1 for each
point shown. For example, you get Rs 6 if the die shows a 6. How much
will you get on average per roll if you play this game 10000
times?
The brute force approach is to really play it 10000 times, and
then divide the total amount by 10000. A smarter way is to use
the probability distribution of the die. We know that on
average we get 1 about 1/6 of the times, 2 about 1/6 of the times
and so on. So on an average we shall get
(1+2+3+4+5+6)/6 = 3.5
per roll. Here the math turned out to be simple. But in more
realistic models the math is often too tough to handle even
with a computer!
-
Simulate probability models: Here also we start with a
model. But instead of dealing with it mathematically, we make the
computer to perform (or simulate) the random experiment following
the model. This is possible because computers can generate random
numbers
 Actually computers generate only pseudo-random
numbers. These are not random but only behave like random
numbers. However, we shall disregard the
difference here. Actually computers generate only pseudo-random
numbers. These are not random but only behave like random
numbers. However, we shall disregard the
difference here. |
Now it is just like the "wait and see" strategy, except that we
do not have to wait long, since computers can perform many
repetitions of a random experiment very fast. Also, no real
monetary loss is involved.
Let us see the third approach in action for the gambling example
before we move on to more serious applications.
values = 1:6 #the possible values
sample(values, 10, replace=T)
This last line asks R to sample 10 numbers from the
vector values. The ``replace=T'' allows
the same number to occur multiple times. Run the last line
repeatedly to see that how the output changes randomly. Now to
solve the problem use
money = sample(values, 10000, replace=T)
avg = mean(money)
avg
This mean is indeed pretty close to the theoretical 3.5. But will
it always be the case? After all it is random game. So it is a
good idea to repeat the above simulation a number of times, say,
100 times. The blue lines below are as before. But these are now
enveloped in some extra lines (shown in bold). This is an example
of a for loop.
avgList = c() #an empty vector
for(i in 1:100) {
money = sample(values, 10000, replace=T)
avg = mean(money)
avgList = c(avgList,avg) #add avg to avgList
}
mean(avgList)
var(avgList)
 |
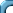 |  |  |
 |
To repeat some commands for a fixed number of times (say 200) in R
you put the commands inside a for loop like this
for(i in 1:200) {
#Your commands
#go here
}
For example, you may try
for(i in 1:5) {
print("hello")
}
|  |
 |  |  |
|
Simulating from a distribution
The fundamental concept behind the above technique is that R can
``roll a die'' or, rather, ``simulate a die''. Statistically
this means generating a random number from among the
values 1,...,6 with probabilities 1/6 each. This is by no
means the only distribution R can generate numbers from. We
can for example, generate a random number from the
Uniform(0,1) distribution. Such a number can take any value
between 0 and 1, each value being equally likely.
Let us generate 100 such numbers.
data = runif(1000, min=0, max=1)
hist(data)
R has many probability distributions built into it, and it has
ready-made functions (like runif) to generate data from
them. The functions all share a common naming pattern. They start
with the letter `r' followed by an acronym for the distribution.
Here are some examples.
data = rnorm(1000, mean=1, sd=2) #Normal(1,2)
hist(data)
data = rpois(1000, lambda=1) #Poisson(1)
hist(data)
data = rexp(1000, rate=2) #Exponential with mean 1/2
hist(data)
Two realistic examples
An imperfect particle counter
A huge radio active source emits particles at random time points. A
particle
detector is detecting them. However, the detector is imperfect
and gets ``jammed'' for 1 sec every time a particle hits it. No
further particles can be detected within this period. After the 1
sec is over, the detector again starts functioning normally. We
want to
know the fraction of particles that the detector is
missing. Since this fraction may depend on the rate at which the
source emits particles we want to get a plot like the following:
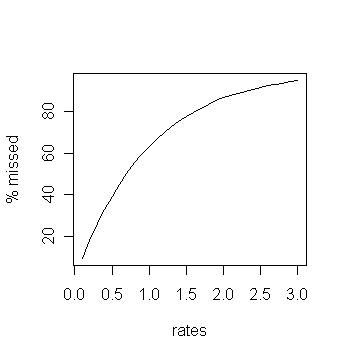 |
| Want a plot like this |
Random emission of particles from a huge radio active source is
well-studied process. A popular model is to assume that the time
gaps between successive emissions are independent and have
Exponential distributions.
gaps = rexp(999,rate=1)
The question now is to determine which of the particles will
fail to be detected. A particle is missed if it comes within 1
sec of its predecessor. So the number of missed particles is
precisely the number of gaps that are below 1.
miss = sum(gaps < 1)
miss
Now we want to repeat this experiment a number of times, say 50
times.
missList = c()
for(i in 1:50) {
gaps = rexp(999,rate=1)
miss = sum(gaps < 1)
missList = c(missList,miss)
}
mean(missList)
var(missList)
All these are done for rate = 1. For other rates we need to repeat
the entire process afresh. So we shall use yet another
for loop.
rates = seq(0.1,3,0.1)
mnList = c()
vrList = c()
for(lambda in rates) {
missList = c()
for(i in 1:50) {
gaps = rexp(1000,rate=lambda)
miss = sum(gaps < 1)
missList = c(missList,miss)
}
mn = mean(missList)
vr = var(missList)
mnList = c(mnList,mn)
vrList = c(vrList, vr)
}
Now we can finally make the plot.
plot(rates,mnList/10,ty="l",ylab="% missed")
We shall throw in two error bars for a good measure.
up = mnList + 2*sqrt(vrList)
lo = mnList - 2*sqrt(vrList)
lines(rates,up/10,col="red")
lines(rates,lo/10,col="blue")
| Exercise: (This exercise is difficult)
We can estimate the actual rate from the hitting times of the
particles if the counter were perfect. The formula is to take the
reciprocal of the average of the gaps. But if we apply the same
formula for the imperfect counter, then we may get a wrong estimate
of the true rate. We want to make a correction
graph that may be used to correct the under estimate.
Explain why the following R program achieves this. You will need
to look up the online help of cumsum and diff.
rates = seq(0.1,3,0.1)
avgUnderEst = c()
for(lambda in rates) {
underEst = c()
for(i in 1:50) {
gaps = rexp(1000,rate=lambda)
hits = cumsum(gaps)
obsHits = c(hits[1], hits[gaps>1])
obsGaps = diff(obsHits)
underEst = c(underEst,1/mean(obsGaps))
}
avgUnderEst = c(avgUnderEst,mean(underEst))
}
plot(avgUnderEst,rates,ty="l")
Can you interpret the plot?
|
Simulating a galaxy
A galaxy is not a single homogeneous body and so it has different
mass densities at different points. When we say that the mass
density of a galaxy at a point (x,y,z) is f(x,y,z)
we mean that for any region R in that galaxy the chance of
detecting a body is
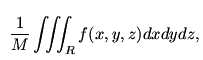
where M is the total mass of the galaxy.
There are various models that specify different forms for
f. Many of these consider the galaxy to be confined in
some 2D plane, so that the density may be written as
f(x,y). The typical process for simulating a galaxy has
two components:
-
First
we simulate the initial state of the galaxy from some probability model.
- Next, we let it evolve deterministically following
appropriate laws of motion.
Such simulations are extremely computation intensive. We
shall therefore confine ourselves to a very simple situation.
To keep life
simple we shall assume that the x and
y-coordinates of the stars are independent normal random
variables. We shall work with just 500 stars of equal masses. (It
is easy to allow random masses, but in our simple
implementation the simulation would then take awfully long to
run!) Each star initially has tangential velocity proportional to
the distance from the center.
x = rnorm(500,sd=4)
y = rnorm(500,sd=4)
vx = -0.5*y
vy = 0.5*x
Let us plot the initial stage of our galaxy.
oldpar = par(bg="black") #we want black background
plot(x,y,pch=".",col="white") #and white stars
par(oldpar) #restore the default (black on white)
This does not look much like a galaxy. We have to let it
evolve over time. This basically means solving an n-body
problem numerically. A popular method is the Hut-Barnes
algorithm. But we shall apply Newton's laws using R in a brute
force (and slow) way. The commands are in the script file
newton.r.
source("newton.r")
Here we shall perform the simulation for 100 time steps, updating
the picture of the galaxy every 10 steps. With only 500 points it
is difficult to see much pattern, but still one may see spiral
tentacles forming.
The simulation here is admittedly very crude and inefficient. If
you are interested, you may see more realistic examples (with
references) at this
website.
Bootstrapping
So far we are simulating from completely specified
distributions. But suppose that someone gives some data and asks
us to simulate from whatever is the distribution of the
data. Since we do not have the distribution itself we cannot
apply the above methods directly.
However, as it happens, there is an approximate way out that is
simple to implement in R.
Let us first see it using an artificial example.
Suppose someone has some data that was originally generated from
N(0,1) distribution.
origData = rnorm(1000,mean=0,sd=1)
Then we shall simply resample from this data set.
newData = sample(origData,500, replace=T)
This is called (one form of) bootstrapping.
It is instructive to compare this with the true distribution.
hist(newData,prob=T)
x = seq(-3,3,0.1)
lines(x,dnorm(x))
Now let us apply this on some real data.
hip = read.table("HIP.dat", header = TRUE, fill = TRUE)
attach(hip)
We shall consider the Vmag variable. We want to
generate 10 new values from its distribution (which is unknown to
us).
newVmag = sample(Vmag,10,replace=T)
Well, that was easily done, but the question is
What in the universe does this newVmag mean?
Does this mean we have generated 10 new galaxies? How can that be
when we are just re-using the old values?
Common sense, please!
First let us beware of the following mistaken notion.
 |
 |  |  |
 |
Mistaken
notion: Bootstrapping is a way to increase the
sample size without any extra real sampling.
|  |
 |  |  |
|
If that were true, you could just keep on
generating further samples from existing data and get away by
pretending that they are new galaxies. Well, common sense
tells us that this cannot be true.
 |
 |  |  |
 |
Great
lesson: Never place statistics above common sense.
|  |
 |  |  |
|
By repeatedly picking from the already available sample we are
not adding anything to the information. In particular, if the
original sample presents a biased view of the underlying
population,
then none of the resamples can cure that distortion.
Why then should we do bootstrapping at all?
The next example explains why this is useful.
Astronomical data sets are often
riddled with outliers (values that are far from the rest). To get
rid of such outliers one sometimes ignores a few of the extreme
values. One such method is the trimmed mean.
x = c(1,2,3,4,5,6,100,7,8,9)
mean(x)
mean(x,trim=0.1) #ignore the extreme 10% points in BOTH ends
mean(x[x!=1 & x!=100])
We shall compute 10%-trimmed mean of Vmag
from the Hipparcos data set.
hip = read.table("HIP.dat", header = TRUE, fill = TRUE)
attach(hip)
mean(Vmag,trim=0.1)
We want to estimate the standard error of this estimate. For
ordinary mean we had a simple formula, but unfortunately such a
simple formula does not exist for the trimmed mean. So we shall
use bootstrap here as follows. We shall generate 100 resamples
each of same size as the original sample.
trmean = c()
for(i in 1:100) {
resamp = sample(Vmag,length(Vmag),replace=T)
trmean = c(trmean,mean(resamp,trim=0.1))
}
sd(trmean)
 |
 |  |  |
 |
Bootstrapping is such a popular statistical technique that R has
a package called boot to perform bootstrapping.
|  |
 |  |  |
|
Permutation test
Here is a variation of the same theme: resampling
i.e., randomly sampling from the original sample and
pretending that it is a new sample.
The situation that we are discussing next involves two samples,
say, the Hyades and the non-Hyades stars. Suppose that we want to
see if the their medians colors are different or not. (By this,
as usual, we
mean to test if the medians of the distributions underlying
the two samples are same or not.) Here we shall assume that the
shapes of the two distributions are the same.
Since the samples are pretty
large, it is reasonable to look at the differences of the medians
of the two samples.
source("hyad.r")
colH = B.V[HyadFilter]
colnH = B.V[!HyadFilter & !is.na(B.V)]
m = length(colH)
n = length(colnH)
median(colH)-median(colnH)
If the population medians are the same, then we should expect
this difference to be near zero, as well. So if this difference
is too large, then we have reason to suspect that the underlying
distributions have different medians. The question therefore is
how large is ``too large"?
Sometimes statistical theory dictates a rule to decide this. But
more often there is no theory to help. Then permutation
tests provide an easy way out.
Imagine first what happens if the medians are the same. Then the
two underlying distributions are actually the same (since their
shapes already match by assumption). So the two samples together
is basically like a single large sample from this common
distribution. If this is the case, calling a Hyades star as
non-Hyades (or vice versa) would really not make any
difference.
So we shall mix up all stars together, and then pick any m
of them and call them Hyades, while the remaining n would
be labeled nonHyades. This should be as good as the original
sample if the two distributions are indeed the same.
pool = c(colH,colnH)
mixed = sample(pool) # sample generates a random permutation
newH = mixed[1:m]
newnH = mixed[(m+1):n]
median(newH) - median(newnH)
If the two distributions were really the same then this new
difference should be close to the difference based on the
original data.
Of course, this apparently takes us back to the original
question: how close is ``close enough"? But now it is easier to
answer since we can repeat this random mixing many times.
pool = c(colH,colnH)
d = c()
for(i in 1:1000) {
mixed = sample(pool)
newH = mixed[1:m]
newnH = mixed[(m+1):n]
newDiff = median(newH) - median(newnH)
d = c(d,newDiff)
}
Now that we have 1000 values of how the typical difference should
look like if the distributions were the same, we can see where
our original difference lies w.r.t. these values. First let us
make a histogram of the typical values:
hist(d)
Well, the original value seems quite far from the range of
typical values, right?
One way to make this idea precise is
to compute the p-value, which is defined as the
chance of observing a difference even more extreme than the
original value. Here ``more extreme" means ``larger in absolute
value".
orig = median(colH)-median(colnH)
sum(abs(d)>=abs(orig))/1000
A p-value smaller than 0.05, say, strongly indicates that
the medians of the distributions are not the same.
 Actually computers generate only pseudo-random
numbers. These are not random but only behave like random
numbers. However, we shall disregard the
difference here.
Actually computers generate only pseudo-random
numbers. These are not random but only behave like random
numbers. However, we shall disregard the
difference here.